Afdeling Radiologie | LUMC
Als afdeling van een Universitair Medisch Centrum richt de radiologie zich vooral op onderzoeken en behandelingen waarvoor extra (wetenschappelijke) kennis nodig is, vergeleken met algemene ziekenhuizen.
In het LUMC doet de afdeling vooral onderzoek naar en behandeling van:
- Hart- en vaatziekten
- Aandoeningen van het hoofd en de hals
- Bottumoren
- Schildklieraandoeningen en neuro-endocriene tumoren (dit zijn tumoren die ontstaan uit cellen die hormonen produceren).
Als afdeling van een Universitair Medisch Centrum richt de radiologie zich vooral op onderzoeken en behandelingen waarvoor extra (wetenschappelijke) kennis nodig is, vergeleken met algemene ziekenhuizen.
In het LUMC doet de afdeling vooral onderzoek naar en behandeling van:
- Hart- en vaatziekten
- Aandoeningen van het hoofd en de hals
- Bottumoren
- Schildklieraandoeningen en neuro-endocriene tumoren (dit zijn tumoren die ontstaan uit cellen die hormonen produceren).
De afdeling Radiologie bestaat uit Algemene Radiologie, Nucleaire Geneeskunde, Laboratorium voor Klinische en Experimentele Beeldverwerking (LKEB), het Gortercentrum en moleculaire beeldvorming. Daarnaast werken wij samen met andere specialismen en verschillende wetenschappelijke instellingen.
Wie zijn wij en wat doen we?
Als onderdeel van een universitair ziekenhuis, richten wij ons vooral op speciale takken van de zorg; de zogeheten "topklinische en topreferente zorg". Wij doen zowel beeldvormende onderzoeken als behandelingen en analyseren verschillende ziekteprocessen..
Er worden jaarlijks ca. 180.000 procedures uitgevoerd op verzoek van medisch specialisten (w.o. huisartsen). Naast de diagnostiek worden er op de verpleegafdeling van de Nucleaire Geneeskunde per jaar ongeveer 250 patiënten behandeld. Het LUMC is daarnaast één van de 10 traumacentra in Nederland voor de opvang en behandeling van complexe ongevalpatiënten. De afdeling Radiologie heeft een taak in het snel diagnose stellen van trauma's (ongevallen).
Meedoen aan onderzoek
Patiëntgebonden wetenschappelijk onderzoek
Wanneer u zich als patiënt laat onderzoeken of behandelen door het LUMC kan het gebeuren, dat u gevraagd wordt om aan wetenschappelijk onderzoek deel te nemen.
Patiëntgebonden wetenschappelijk onderzoek is van groot belang voor het ontwikkelen van nieuwe onderzoeken of behandelingen. U bent volledig vrij in uw keuze en uw besluit heeft geen effect op uw behandeling.
…Patiëntgebonden wetenschappelijk onderzoek
Wanneer u zich als patiënt laat onderzoeken of behandelen door het LUMC kan het gebeuren, dat u gevraagd wordt om aan wetenschappelijk onderzoek deel te nemen.
Patiëntgebonden wetenschappelijk onderzoek is van groot belang voor het ontwikkelen van nieuwe onderzoeken of behandelingen. U bent volledig vrij in uw keuze en uw besluit heeft geen effect op uw behandeling.
Gezonde vrijwilligers
Indien u als gezonde vrijwilliger deel wilt nemen aan een wetenschappelijk onderzoek kunt u zich ook opgeven.
Zie voor meer informatie over proefpersonen / wetenschappelijk onderzoek binnen het LUMC: Bontiusstichting.
Doorverwijzen, aanvragen en uitslagen
Binnen de afdeling Radiologie worden zowel beeldvormende diagnostiek als door beeldvorming ondersteunde diagnostische en therapeutische interventies verricht. Er worden jaarlijks ca. 180.000 procedures uitgevoerd op verzoek van medisch specialisten.
Polikliniek
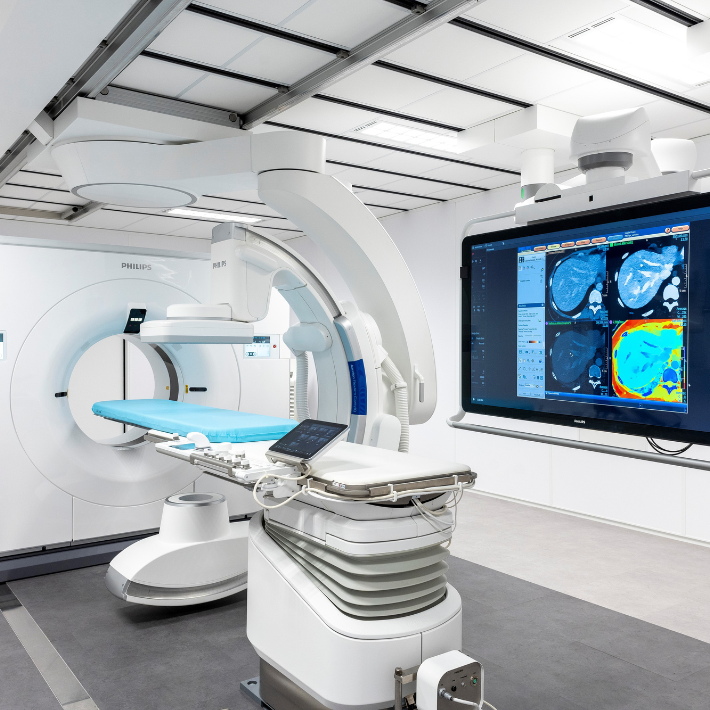
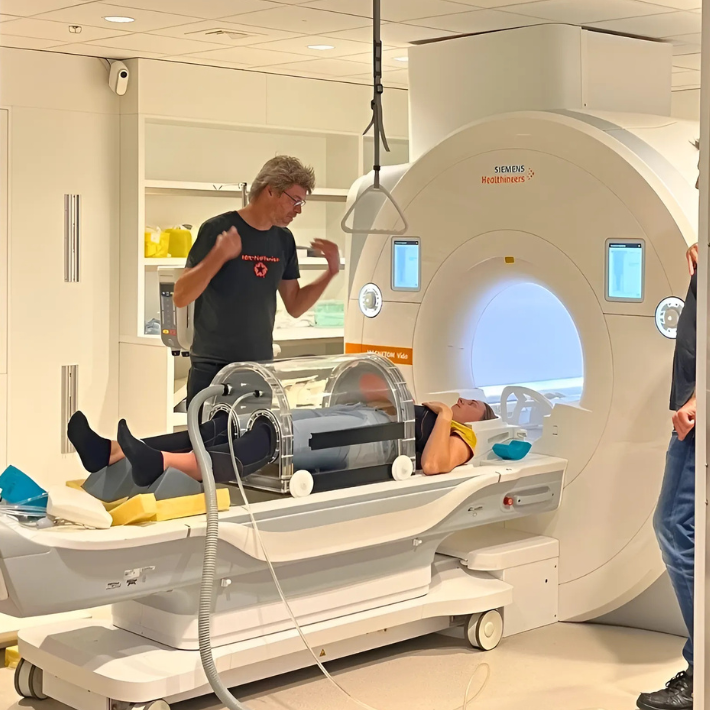
Als u een röntgenfoto, echografie of MRI-scan moet laten maken, kunt u terecht op de polikliniek Radiologie. Hier verrichten wij onderzoeken en stellen wij diagnoses door middel van stralen of golven. Zo kunnen wij de aard en plaats van een ziekte, letsel of aandoening achterhalen.
De afdeling Radiologie bestaat uit Conventionele Radiologie, Mammografie, CT, MRI, Interventie Radiologie, Echo en Nucleaire Geneeskunde.
Veiligheid en kwaliteit
Het LUMC leidt o.a. artsen op tot radioloog / nucleair geneeskundige, samen met ziekenhuizen in de omgeving. Daarom kunt u ook onderzocht en/of behandeld worden door artsen in opleiding. De kwaliteit van deze opleiding wordt gegarandeerd door de Koninklijke Nederlandse Specialisten Vereniging.
Het onderwijs richt zich op het opleiden en bij- en nascholen van verschillende specialisten zoals radiologen, nucleair geneeskundigen, klinisch fysici, (bio)medische studenten en studenten Medisch Beeldvormende en Radiotherapeutische Technieken (HBO-MBRT).
…Het LUMC leidt o.a. artsen op tot radioloog / nucleair geneeskundige, samen met ziekenhuizen in de omgeving. Daarom kunt u ook onderzocht en/of behandeld worden door artsen in opleiding. De kwaliteit van deze opleiding wordt gegarandeerd door de Koninklijke Nederlandse Specialisten Vereniging.
Het onderwijs richt zich op het opleiden en bij- en nascholen van verschillende specialisten zoals radiologen, nucleair geneeskundigen, klinisch fysici, (bio)medische studenten en studenten Medisch Beeldvormende en Radiotherapeutische Technieken (HBO-MBRT).
Wij bieden ondersteuning voor iedere carrièrefase in de radiologie. Zo geven wij informatie aan scholieren voor hun eventuele beroepskeuze in de radiologie. We verzorgen hoorcolleges over beeldvormende technieken op HBO en universitair niveau. Daarnaast bieden wij mogelijkheden tot het volgen van zowel technische als klinische stages, leiden we op tot radioloog, nucleair geneeskundige of klinisch fysicus, en geven we bij- en na-scholing aan specialisten.
Ouderdag
Elk jaar organiseren eerstejaars studenten Biomedische Wetenschappen en Geneeskunde van de Medische Faculteit Leidse Studenten een ouderdag, waarop ouders kunnen zien waar hun kinderen mee bezig zijn. Hieraan neemt de afdeling Radiologie deel, d.m.v. een presentatie, gehouden door AIO’ s, waarin de studenten en hun ouders wordt uitgelegd wat de radiologie inhoudt. Daarnaast worden ze over de afdeling rondgeleid.
…Ouderdag
Elk jaar organiseren eerstejaars studenten Biomedische Wetenschappen en Geneeskunde van de Medische Faculteit Leidse Studenten een ouderdag, waarop ouders kunnen zien waar hun kinderen mee bezig zijn. Hieraan neemt de afdeling Radiologie deel, d.m.v. een presentatie, gehouden door AIO’ s, waarin de studenten en hun ouders wordt uitgelegd wat de radiologie inhoudt. Daarnaast worden ze over de afdeling rondgeleid.
Medische Carrière Dag
Op deze dag, georganiseerd door de Medische Faculteit Leidse Studenten, krijgen geneeskunde studenten en co-assistenten de mogelijkheid zich te oriënteren op de carrièremogelijkheden na het artsexamen. De dag bestaat uit een aantal parallelsessies, een informatiemarkt en een borrel. De afdeling Radiologie heeft een stand op de informatiemarkt.
Opleidingen
The Department of Radiology has a longstanding tradition in scientific research at the interface of technological innovations and healthcare. Reflecting this tradition, our research program is organized in themes with a technological focus on the one hand, and themes with an organ-centered biomedical focus on the other hand.
Research programs
Our aim is to develop imaging technologies that can be used in patients for diagnostic purposes or as instruments to increase our understanding of diseases. Consequently, research in our technological themes is tuned to the needs in our biomedical themes. In practice, our research is characterized by close interdisciplinary interactions across the various themes and with partners from outside our department.
The themes of our research program are embedded in the following research groups:
…Our aim is to develop imaging technologies that can be used in patients for diagnostic purposes or as instruments to increase our understanding of diseases. Consequently, research in our technological themes is tuned to the needs in our biomedical themes. In practice, our research is characterized by close interdisciplinary interactions across the various themes and with partners from outside our department.
The themes of our research program are embedded in the following research groups:
Imaging technology based lines:
- Automated image processing and quantification (coordinator Prof. Dr. Ir. B.J. Lelieveldt
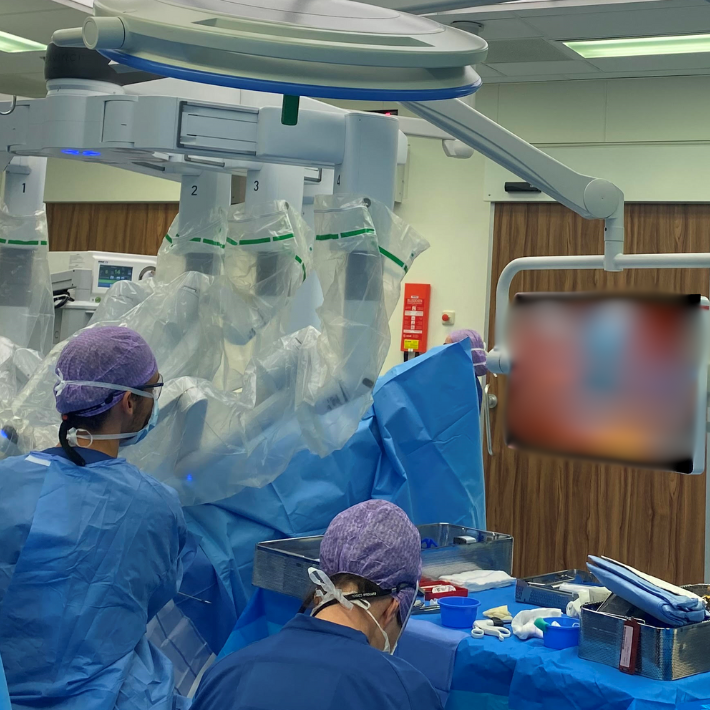
- Interventional radiology (coordinator dr. M.C. Burgmans)
- Interventional/intraoperative molecular imaging (coordinator Prof. Dr. F.W.B. van Leeuwen)
- MRI (coordinator Prof. Dr. Ir. M.J.P van Osch)
- Nanomedicine and Molecular Imaging (coordinator Dr. L.J. Cruz)
- Nuclear Medicine (coordinator Prof. Dr. L.F. de Geus-Oei)
Organ based lines:
- Abdominal (coordinator Dr. M.N.J.M. Wasser)
- Breast (coordinator Drs. E.M.C Voormolen)
- Cardiovascular & thoracic (coordinator Prof. Dr. H.J. Lamb)
- Musculoskeletal (coordinator Dr. K. van Langevelde)
- Neuroradiology (coordinator Dr. J.H.J. M. de Bresser)
Research organisation
The Group Leader Board (GLB) is responsible for the organizational aspects of our research programmes. All project leaders of our research lines and the manager of the department are members of the GLB. Two members of the GLB are representatives, one with a clinical and one with a technological background. They are part of the department and report to the chair of the Department of Radiology.
The Scientific Committee (Wetenschapscommissie) is responsible for assessing the quality and feasibility of new research proposals from within and outside the department, keeping track of accepted projects, and distributing time slots for research on our imaging systems. The chair of this committee reports to the MTR.
The Radiology Consultancy Center (RCC) of our department provides services for large-scale patient studies in terms of study design, data analysis, and data storage. The chair of the RCC reports to the MTR.
Contactgegevens
Postadres
Postzone
Postbus 9600
2300 RC Leiden
Bezoekadres
Secretariaat:
Locatie: C3-S
Telefoonnummer: 071 526 3474
Informatie voor de patiënt
Buiten kantooruren kunt u de dienstdoende arts / radioloog bereiken via de telefooncentrale van het LUMC Telefoonnummer 071 526 91 11.
Binnen kantooruren (08.00-16.30 uur) kunt u als patiënt de afdeling Radiologie telefonisch bereiken via onze receptionisten.
…Postadres
Postzone
Postbus 9600
2300 RC Leiden
Bezoekadres
Secretariaat:
Locatie: C3-S
Telefoonnummer: 071 526 3474
Informatie voor de patiënt
Buiten kantooruren kunt u de dienstdoende arts / radioloog bereiken via de telefooncentrale van het LUMC Telefoonnummer 071 526 91 11.
Binnen kantooruren (08.00-16.30 uur) kunt u als patiënt de afdeling Radiologie telefonisch bereiken via onze receptionisten.
- Echografie, röntgenfoto's van het skelet, maag-darm, nieren, hart of longen, mammografie en interventie-onderzoeken:
Telefoonnummer 071 526 20 32
e-mail: Radiologie-afsprakenbureau@lumc.nl - CT- en MRI-scan:
Telefoonnummer 071 526 18 40
e-mail: RadiologieafsprakenCT-MRI@lumc.nl
- Nucleaire Geneeskunde, tweede verdieping, receptie C2-P.
Telefoonnummer 071 526 34 80
e-mail: Nuge_receptie@lumc.nl
Let op: voor deze locatie gelden afwijkende kantooruren (08.30-10.30 uur) en (13.30-16.00 uur)
Informatie voor verwijzers
Binnen kantooruren (08.00-16.30 uur) kunt u de administratie van de afdeling Radiologie telefonisch bereiken via Telefoonnummer 071 526 20 32.
Telefoonnummer voor afspraken over onderstaande onderzoeken of behandelingen:
071 526 24 10
- Angio- en neuro-interventies
071 526 20 32
- Echografie
- Maag-darm onderzoek
- Mammografie
- Orthopaedie/ musculo skelettaal onderzoek
- Thoraxopnamen
- en ander conventioneel onderzoek aan te vragen door de huisartsen
071 526 18 40
- CT onderzoek
- MRI onderzoek
071 526 34 80
- Nucleair geneeskundig onderzoek of behandeling
Beeldmateriaal kan tijdens kantooruren worden opgevraagd, per telefoon op 071 526 20 32 of per email aan lumc-beeldenuitwisseling@lumc.nl.
U kunt met technische/praktische vragen of opmerkingen ten aanzien van digitale beelduitwisseling contact met ons opnemen door een email te sturen aan lumc-beeldenuitwisseling@lumc.nl of tijdens kantooruren te bellen met telefoonnummer: 071 526 34 97.
Alleen in spoedeisende gevallen kunt u buiten kantooruren contact opnemen met de dienstdoende laborant Radiologie, te bereiken via de telefooncentrale van het LUMC: 071 526 91 11.
Contact over Onderwijs en Research
Veelgestelde vragen
Research Consultancy Center
Het Research Consultancy Center (RCC) van de afdeling Radiologie van het LUMC biedt professionele ondersteuning bij zowel het opzetten als uitvoeren van radiologisch georiënteerd, mens-gebonden onderzoek op onze afdeling. Doel is om een positieve bijdrage te leveren aan het juiste design van onderzoek en aan de kwaliteit en integriteit van data, beeldbewerking en statistische analyses.
Het team bestaat uit een groep biomedische medewerkers met specifieke expertise, allen getraind op het gebied van Good Research Practice. Het RCC is het startpunt voor elk wetenschappelijk onderzoek op de afdeling Radiologie van het LUMC en is ook bij uitstek een geschikte contractpartner voor commercieel gefinancierd onderzoek.
&width=180&height=180)
&width=180&height=180)
&width=180&height=180)